If you would like to discuss possible master thesis topics, please have a look at my staff page and send me an email. Also make sure to check out postdoc Ove Øyås' offerings; we collaborate closely.
I am passionate about using mathematics to integrate biological knowledge and data, identifying patterns and contrasts, making comparisons and making real-world interpretations of the results. The production biology of salmon is my current focus, though I also co-supervise on medical genetics, studying the clinical importance of regulatory variation in DNA and micro-RNA. I believe in combining approaches ranging from broad statistical descriptions to more mechanistic mathematical descriptions of physiology. The value of modeling is purposeful simplification of reality, generalizing insights from the model to a broader class of systems.
Systems biology means translating what we know about biology into numbers and equations, so that we can use computer simulations and the rules of mathematics to deduce new insights and hypothesis. The Digital Salmon project lays the foundations for "digital production biology", starting with the challenge of sustainably feeding farmed salmon. The models we build will eventually also serve as a framework for comparison with and among wild populations, documenting salmonid biodiversity and aiding ecological management.
DigiSal is part of Digital Life Norway, the Research Council's biggest program for systems biology. The common theme is life, technology and value generation – by applying mathematics.
If you love biochemistry, math and computer programming.
Everything you are is made from stuff you ate. The building up and breaking down of living material happens through chemical reactions catalyzed by enzymes coded for by the genes in your DNA. The same is true for every organism.
The biochemical reaction network covers the wall of my office. It is beautiful:
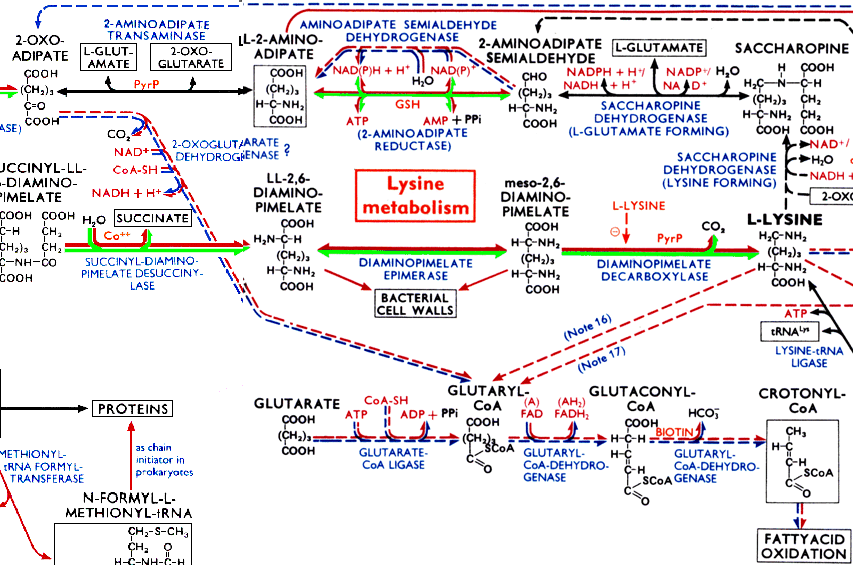
The full chart is huge! Have a look at this interactive biochemical pathway map (c) Hoffman-La Roche Ltd.
The first models in the Digital Salmon translate salmon's metabolic reaction network to mathematics and computer programming.
If you love genetics and bioinformatics.
Simon Rayner studies the importance of the small RNAs that do not code for proteins, but rather modify the expression and usage of other genes. In particular, his group uses computer programming to search for genetic variants associated with disease in large databases of human genomes. Here's a poster illustrating their work, and a summary:
miRNAs are short non-coding RNAs that, together with transcription factors, play a central role in gene regulation. miRNAs are produced from the stem loop of pre-cursor elements referred to as pre-miRNAs and are found in diverse eukaryotic lineages as well as some dsDNA viruses. miRNAs are typically 22nt in length but functional forms as short as 16nt and as long as 30nt have been identified. The most commonly studied function is the process by which they bind to the 3’UTR of target mRNAs, leading to translational repression or mRNA decay.
The majority of miRNA studies focus on identifying their functional roles in disease and developmental processes. These functional studies have been particularly fruitful, for example, associating individual miRNAs with key developmental roles or disease. In the latter case, these miRNAs can often be used as biomarkers for a specific disease. In the most common approach, “miRNA association studies” are performed using microarray or NGS platforms to compare two conditions (e.g. healthy versus cancer) and identify miRNAs that have statistically significant differences in expression levels. The mRNA targets of these miRNAs are determined using computational prediction tools and functionally interesting ones are experimentally verified. Follow up studies in animal models may also be performed to provide further experimental support.
However, the targeting process is still poorly understood and computational approaches demonstrate mixed performance. We recently applied a deep learning approach to the problem and, while our trained model outperforms other software tools, we still lack detailed understanding of the mechanisms driving the targeting process. Therefore, our next step is to experimentally generate new positive and negative data and use investigate how they impact the model in terms of specific predictions of miRNA-3’UTR binding patterns and general performance.
If you love experimenting with computer code.
Postdoc Ove Øyås and I collaborate with William Harcombe's group at the University of Minnesota. Their simulation framework for Computation of Microbial Ecosystems in Time and Space (COMETS) opens possibilities for many exciting computer simulation experiments.
I have broad interests and varied experience in biological data analysis and programming. If you have contact with a research group or organization with biological data that they want to analyse, we can discuss co-supervision.